Learned Sensing: Jointly optimized microscope hardware for accurate image classification
Biomedical Optics Express (2019)
Alex Muthumbi1,+,Amey Chaware2,+, Kanghyun Kim2, Kevin Zhou3, Pavan Chandra Konda3, Richard Chen4, Benjamin Judkewitz5, Andreas Erdmann6,1, Barbara Kappes7, Roarke Horstmeyer2,3
1: School of Advanced Optical Technologies, Friedrich-Alexander University, Erlangen, Germany.
2: Department of Electrical and Computer Engineering, Duke University, Durham NC, USA.
3: Department of Biomedical Engineering, Duke University, Durham NC, USA.
4: Y Combinator Research, San Francisco, CA.
5: NeuroCure Cluster of Excellence, Charite Universitatsmedizin and Humboldt University, Berlin, Germany.
6: Fraunhofer IISB, Erlangen, Germany.
7: Department of Chemical and Biological Engineering, Friedrich-Alexander University, Erlangen, Germany.
+ denotes equally contributing authors
Links to paper, code, and data
Abstract
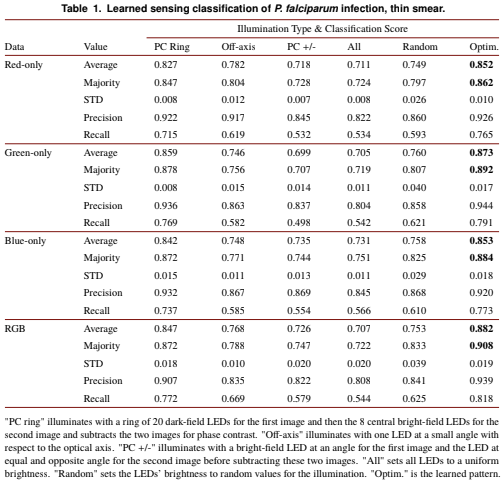
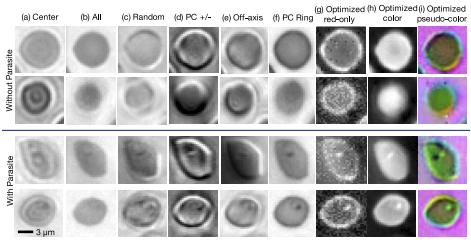
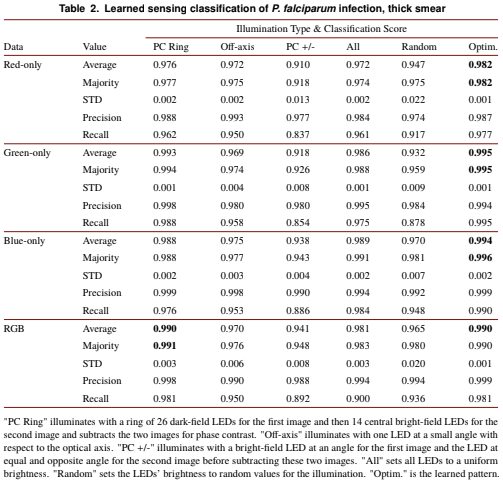
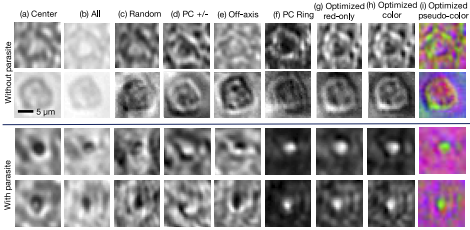
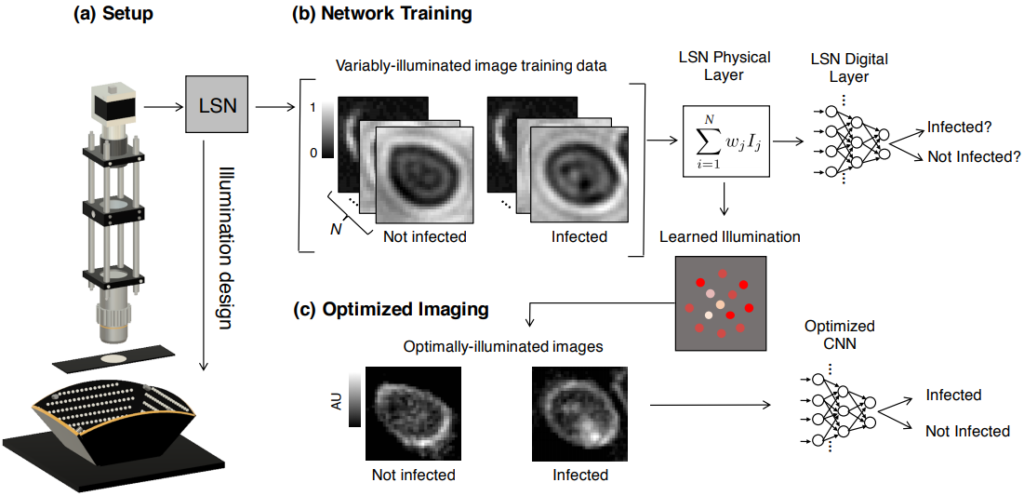
Since its invention, the microscope has been optimized for interpretation by a human observer. With the recent development of deep learning algorithms for automated image analysis, there is now a clear need to re-design the microscope’s hardware for specific interpretation tasks. To increase the speed and accuracy of automated image classification, this work presents a method to co-optimize how a sample is illuminated in a microscope, along with a pipeline to automatically classify the resulting image, using a deep neural network. By adding a ”physical layer” to a deep classification network, we are able to jointly optimize for specific illumination patterns that highlight the most important sample features for the particular learning task at hand, which may not be obvious under standard illumination. We demonstrate how our learned sensing approach for illumination design can automatically identify malaria-infected cells with up to 5-10% greater accuracy than standard and alternative microscope lighting designs. We show that this joint hardware-software design procedure generalizes to offer accurate diagnoses for two different blood smear types, and experimentally show how our new procedure can translate across different experimental setups while maintaining high accuracy.
Our Setup
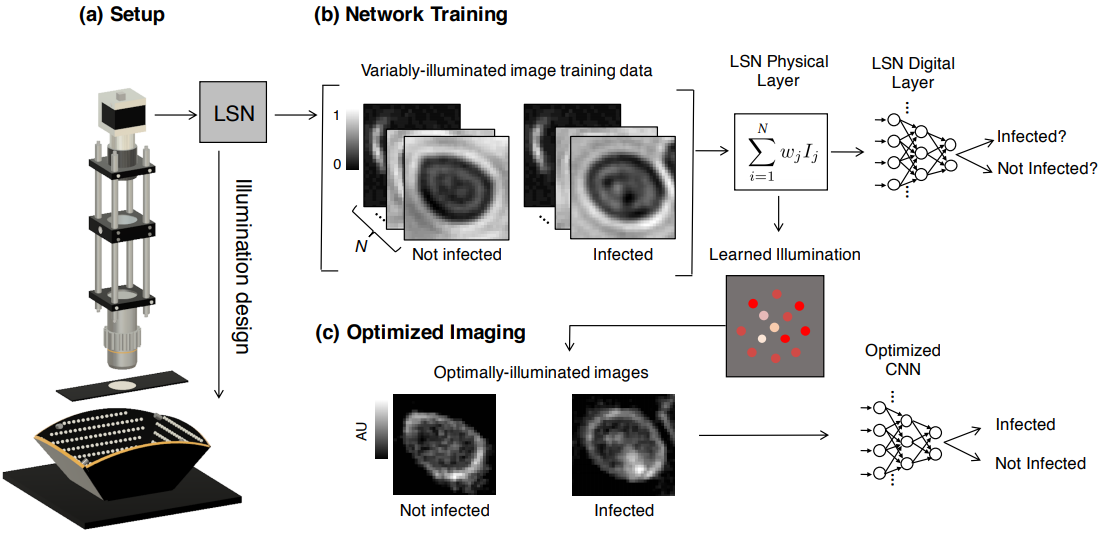
Results